


|
 |
#include <MuScleFitBase.h>
Classes | |
| class | ProbForIntegral |
| Functor used to compute the normalization integral of probability functions. More... | |
Public Member Functions | |
| MuScleFitBase (const edm::ParameterSet &iConfig) | |
| virtual | ~MuScleFitBase () noexcept(false) |
Protected Member Functions | |
| void | clearHistoMap () |
| Clean the histograms map. More... | |
| void | fillHistoMap (TFile *outputFile, unsigned int iLoop) |
| Create the histograms map. More... | |
| void | readProbabilityDistributionsFromFile () |
| Read probability distributions from a local root file. More... | |
| void | writeHistoMap (const unsigned int iLoop) |
| Save the histograms map to file. More... | |
Protected Attributes | |
| int | debug_ |
| std::vector< GenMuonPair > | genMuonPairs_ |
| Stores the genMuon pairs and the motherId prior to the creation of the internal tree. More... | |
| std::map< std::string, Histograms * > | mapHisto_ |
| The map of histograms. More... | |
| std::vector< MuonPair > | muonPairs_ |
| Used to store the muon pairs plus run and event number prior to the creation of the internal tree. More... | |
| std::string | probabilitiesFile_ |
| std::string | probabilitiesFileInPath_ |
| int | theCompressionSettings_ |
| std::vector< TFile * > | theFiles_ |
| The files were the histograms are saved. More... | |
| std::string | theGenInfoRootFileName_ |
| edm::InputTag | theMuonLabel_ |
| int | theMuonType_ |
| std::string | theRootFileName_ |
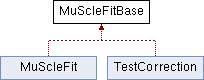
This class is used as a base for MuSlceFit. The reason for putting some of the methods inside this base class is that they are used also by the TestCorrection analyzer.
Definition at line 18 of file MuScleFitBase.h.
|
inline |
Definition at line 20 of file MuScleFitBase.h.
|
inlinevirtualnoexcept |
Definition at line 31 of file MuScleFitBase.h.
|
protected |
Clean the histograms map.
Definition at line 143 of file MuScleFitBase.cc.
References timingPdfMaker::histo, and mapHisto_.
Referenced by MuScleFit::endOfFastLoop().
|
protected |
Create the histograms map.
Definition at line 9 of file MuScleFitBase.cc.
References MuScleFitUtils::debugMassResol_, LogDebug, mapHisto_, B2GTnPMonitor_cfi::maxMass, PV_cfg::maxPt, B2GTnPMonitor_cfi::minMass, download_sqlite_cfg::outputFile, MuScleFitUtils::resfind, and theMuonType_.
Referenced by MuScleFit::startingNewLoop(), and TestCorrection::TestCorrection().
|
protected |
Read probability distributions from a local root file.
Definition at line 159 of file MuScleFitBase.cc.
References gather_cfg::cout, beamvalidation::exit(), geometryDiff::file, GL, MuScleFitUtils::GLNorm, MuScleFitUtils::GLValue, MuScleFitUtils::GLZNorm, MuScleFitUtils::GLZValue, mps_fire::i, ALPAKA_ACCELERATOR_NAMESPACE::ecal::reconstruction::internal::endcap::ix(), ALPAKA_ACCELERATOR_NAMESPACE::ecal::reconstruction::internal::endcap::iy(), MuScleFitUtils::nbins, probabilitiesFile_, probabilitiesFileInPath_, MuScleFitUtils::rapidityBinsForZ_, MuScleFitUtils::resfind, MuScleFitUtils::ResHalfWidth, MuScleFitUtils::ResMaxSigma, MuScleFitUtils::ResMinMass, and theMuonType_.
Referenced by MuScleFit::beginOfJobInConstructor(), and TestCorrection::initialize().
|
protected |
Save the histograms map to file.
Definition at line 150 of file MuScleFitBase.cc.
References timingPdfMaker::histo, mapHisto_, and theFiles_.
Referenced by MuScleFit::endOfFastLoop(), and TestCorrection::~TestCorrection().
|
protected |
Definition at line 53 of file MuScleFitBase.h.
Referenced by MuScleFit::applyBias(), MuScleFit::applySmearing(), MuScleFit::beginOfJobInConstructor(), MuScleFit::checkDeltaR(), MuScleFit::duringFastLoop(), MuScleFit::duringLoop(), MuScleFit::endOfJob(), TestCorrection::fillMuonCollection(), MuScleFit::fillMuonCollection(), MuScleFit::MuScleFit(), MuScleFit::selectMuons(), MuScleFit::startingNewLoop(), and MuScleFit::~MuScleFit().
|
protected |
Stores the genMuon pairs and the motherId prior to the creation of the internal tree.
Definition at line 84 of file MuScleFitBase.h.
Referenced by MuScleFit::selectMuons(), and MuScleFit::~MuScleFit().
|
protected |
The map of histograms.
Definition at line 79 of file MuScleFitBase.h.
Referenced by TestCorrection::analyze(), clearHistoMap(), MuScleFit::duringFastLoop(), MuScleFit::fillComparisonHistograms(), fillHistoMap(), and writeHistoMap().
|
protected |
Used to store the muon pairs plus run and event number prior to the creation of the internal tree.
Definition at line 82 of file MuScleFitBase.h.
Referenced by MuScleFit::selectMuons(), and MuScleFit::~MuScleFit().
|
protected |
Definition at line 45 of file MuScleFitBase.h.
Referenced by readProbabilityDistributionsFromFile().
|
protected |
Definition at line 44 of file MuScleFitBase.h.
Referenced by readProbabilityDistributionsFromFile().
|
protected |
Definition at line 49 of file MuScleFitBase.h.
Referenced by MuScleFit::beginOfJobInConstructor().
|
protected |
The files were the histograms are saved.
Definition at line 76 of file MuScleFitBase.h.
Referenced by MuScleFit::beginOfJobInConstructor(), MuScleFit::endOfFastLoop(), MuScleFit::startingNewLoop(), TestCorrection::TestCorrection(), writeHistoMap(), and TestCorrection::~TestCorrection().
|
protected |
Definition at line 51 of file MuScleFitBase.h.
Referenced by MuScleFit::beginOfJobInConstructor().
|
protected |
Definition at line 48 of file MuScleFitBase.h.
Referenced by MuScleFit::MuScleFit().
|
protected |
Definition at line 47 of file MuScleFitBase.h.
Referenced by TestCorrection::analyze(), fillHistoMap(), MuScleFit::MuScleFit(), readProbabilityDistributionsFromFile(), MuScleFit::selectMuons(), MuScleFit::takeSelectedMuonType(), and MuScleFit::~MuScleFit().
|
protected |
Definition at line 50 of file MuScleFitBase.h.
Referenced by MuScleFit::beginOfJobInConstructor(), and TestCorrection::TestCorrection().
 1.8.14
1.8.14