


|
 |
#include <HIPUserVariablesIORoot.h>
Public Types | |
| typedef std::vector< Alignable * > | Alignables |
Public Member Functions | |
| HIPUserVariablesIORoot () | |
| std::vector < AlignmentUserVariables * > | readHIPUserVariables (const Alignables &alivec, const char *filename, int iter, int &ierr) |
| void | writeHIPUserVariables (const Alignables &alivec, const char *filename, int iter, bool validCheck, int &ierr) |
Private Types | |
| typedef std::map< std::pair < int, int >, int > | treemaptype |
Private Member Functions | |
| int | close (void) |
| void | createBranches (void) |
| create root branches More... | |
| int | findEntry (unsigned int detId, int comp) |
| int | open (const char *filename, int iteration, bool writemode) |
| AlignmentUserVariables * | readOne (Alignable *ali, int &ierr) |
| void | setBranchAddresses (void) |
| set root branches More... | |
| int | writeOne (Alignable *ali) |
Private Attributes | |
| double | AlignableChi2 |
| unsigned int | AlignableNdof |
| unsigned int | Id |
| double | Jtve [nparmax] |
| double | Jtvj [nparmax *(nparmax+1)/2] |
| bool | newopen |
| int | Nhit |
| int | Npare |
| int | Nparj |
| int | ObjId |
| treemaptype | treemap |
Static Private Attributes | |
| static const int | nparmax =6 |
Additional Inherited Members | |
 Protected Member Functions inherited from AlignmentIORootBase Protected Member Functions inherited from AlignmentIORootBase | |
| AlignmentIORootBase () | |
| constructor More... | |
| int | closeRoot (void) |
| close IO More... | |
| int | openRoot (const char *filename, int iteration, bool writemode) |
| open IO More... | |
| int | testFile (const char *filename, const TString &tname) |
| test if file is existing and if so, what the highest iteration is More... | |
| TString | treeName (int iter, const TString &tname) |
| compose tree name More... | |
| virtual | ~AlignmentIORootBase () |
| destructor More... | |
 Protected Member Functions inherited from AlignmentUserVariablesIO Protected Member Functions inherited from AlignmentUserVariablesIO | |
| std::vector < AlignmentUserVariables * > | read (const align::Alignables &alivec, int &ierr) |
| int | write (const align::Alignables &alivec, bool validCheck) |
| virtual | ~AlignmentUserVariablesIO () |
 Protected Attributes inherited from AlignmentIORootBase Protected Attributes inherited from AlignmentIORootBase | |
| bool | bWrite |
| TTree * | tree |
| TString | treename |
| TString | treetxt |
 Static Protected Attributes inherited from AlignmentIORootBase Static Protected Attributes inherited from AlignmentIORootBase | |
| static const int | itermax = 1000 |
| static const int | nParMax = 20 |
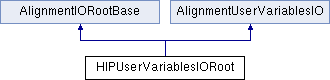
concrete class for ROOT based IO of AlignmentUserVariables
Definition at line 9 of file HIPUserVariablesIORoot.h.
| typedef std::vector<Alignable*> HIPUserVariablesIORoot::Alignables |
Definition at line 15 of file HIPUserVariablesIORoot.h.
|
private |
Definition at line 65 of file HIPUserVariablesIORoot.h.
| HIPUserVariablesIORoot::HIPUserVariablesIORoot | ( | ) |
constructor
Definition at line 16 of file HIPUserVariablesIORoot.cc.
References i, Jtve, Jtvj, nparmax, AlignmentIORootBase::treename, and AlignmentIORootBase::treetxt.
|
inlineprivatevirtual |
close IO
Implements AlignmentUserVariablesIO.
Definition at line 43 of file HIPUserVariablesIORoot.h.
References AlignmentIORootBase::closeRoot().
Referenced by lumiQTWidget.ApplicationWindow::fileQuit(), esMonitoring.AsyncLineReaderMixin::handle_close(), esMonitoring.FDJsonServer::handle_close(), Vispa.Gui.BoxContentDialog.BoxContentDialog::keyPressEvent(), Vispa.Gui.FindDialog.FindDialog::keyPressEvent(), readHIPUserVariables(), and writeHIPUserVariables().
|
privatevirtual |
create root branches
Implements AlignmentIORootBase.
Definition at line 31 of file HIPUserVariablesIORoot.cc.
References AlignableChi2, AlignableNdof, Id, Jtve, Jtvj, Nhit, Npare, Nparj, ObjId, and AlignmentIORootBase::tree.
|
private |
Definition at line 65 of file HIPUserVariablesIORoot.cc.
References ev, Id, newopen, ObjId, query::result, AlignmentIORootBase::tree, and treemap.
Referenced by QGLikelihoodCalculator.QGLikelihoodCalculator::computeQGLikelihood(), and readOne().
|
inlineprivatevirtual |
open IO
Implements AlignmentUserVariablesIO.
Definition at line 39 of file HIPUserVariablesIORoot.h.
References newopen, and AlignmentIORootBase::openRoot().
Referenced by Vispa.Plugins.ConfigEditor.ConfigEditorTabController.ConfigEditorTabController::importConfig(), readHIPUserVariables(), and writeHIPUserVariables().
| std::vector< AlignmentUserVariables * > HIPUserVariablesIORoot::readHIPUserVariables | ( | const Alignables & | alivec, |
| const char * | filename, | ||
| int | iter, | ||
| int & | ierr | ||
| ) |
read user variables
Definition at line 190 of file HIPUserVariablesIORoot.cc.
References close(), open(), AlignmentUserVariablesIO::read(), and query::result.
Referenced by HIPAlignmentAlgorithm::collector().
|
privatevirtual |
read AlignmentParameters of one Alignable
Implements AlignmentUserVariablesIO.
Definition at line 134 of file HIPUserVariablesIORoot.cc.
References HIPUserVariables::alichi2, AlignableChi2, AlignableNdof, Alignable::alignableObjectId(), HIPUserVariables::alindof, cuy::col, KineDebug3::count(), findEntry(), Alignable::id(), HIPUserVariables::jtve, Jtve, HIPUserVariables::jtvj, Jtvj, HIPUserVariables::nhit, Nhit, np, Npare, and AlignmentIORootBase::tree.
|
privatevirtual |
set root branches
Implements AlignmentIORootBase.
Definition at line 47 of file HIPUserVariablesIORoot.cc.
References AlignableChi2, AlignableNdof, Id, Jtve, Jtvj, Nhit, Npare, Nparj, ObjId, and AlignmentIORootBase::tree.
| void HIPUserVariablesIORoot::writeHIPUserVariables | ( | const Alignables & | alivec, |
| const char * | filename, | ||
| int | iter, | ||
| bool | validCheck, | ||
| int & | ierr | ||
| ) |
write user variables
Definition at line 174 of file HIPUserVariablesIORoot.cc.
References close(), open(), and AlignmentUserVariablesIO::write().
Referenced by HIPAlignmentAlgorithm::terminate().
|
privatevirtual |
write AlignmentParameters of one Alignable
Implements AlignmentUserVariablesIO.
Definition at line 94 of file HIPUserVariablesIORoot.cc.
References HIPUserVariables::alichi2, AlignableChi2, AlignableNdof, Alignable::alignableObjectId(), Alignable::alignmentParameters(), HIPUserVariables::alindof, cuy::col, KineDebug3::count(), Id, Alignable::id(), HIPUserVariables::jtve, Jtve, HIPUserVariables::jtvj, Jtvj, HIPUserVariables::nhit, Nhit, np, Npare, Nparj, ObjId, AlignmentIORootBase::tree, and AlignmentParameters::userVariables().
|
private |
Definition at line 61 of file HIPUserVariablesIORoot.h.
Referenced by createBranches(), readOne(), setBranchAddresses(), and writeOne().
|
private |
Definition at line 62 of file HIPUserVariablesIORoot.h.
Referenced by createBranches(), readOne(), setBranchAddresses(), and writeOne().
|
private |
Definition at line 57 of file HIPUserVariablesIORoot.h.
Referenced by createBranches(), findEntry(), setBranchAddresses(), and writeOne().
|
private |
Definition at line 60 of file HIPUserVariablesIORoot.h.
Referenced by createBranches(), HIPUserVariablesIORoot(), readOne(), setBranchAddresses(), and writeOne().
Definition at line 59 of file HIPUserVariablesIORoot.h.
Referenced by createBranches(), HIPUserVariablesIORoot(), readOne(), setBranchAddresses(), and writeOne().
|
private |
Definition at line 64 of file HIPUserVariablesIORoot.h.
Referenced by findEntry(), and open().
|
private |
Definition at line 58 of file HIPUserVariablesIORoot.h.
Referenced by createBranches(), readOne(), setBranchAddresses(), and writeOne().
|
private |
Definition at line 58 of file HIPUserVariablesIORoot.h.
Referenced by createBranches(), readOne(), setBranchAddresses(), and writeOne().
|
private |
Definition at line 58 of file HIPUserVariablesIORoot.h.
Referenced by createBranches(), setBranchAddresses(), and writeOne().
|
staticprivate |
Definition at line 53 of file HIPUserVariablesIORoot.h.
Referenced by HIPUserVariablesIORoot().
|
private |
alignment parameter tree
Definition at line 56 of file HIPUserVariablesIORoot.h.
Referenced by createBranches(), findEntry(), setBranchAddresses(), and writeOne().
|
private |
Definition at line 66 of file HIPUserVariablesIORoot.h.
Referenced by findEntry().
 1.8.5
1.8.5